Sequencher contig
Euphotic zone free DNA (75 to 125 m) contained primarily bacterial and viral sequences, with bacteria dominating samples from the mesopelagic zone (500 to 1,000 m). Viral DNA consisted predominantly of myovirus-like and podovirus-like DNA and contained the highest proportion of unannotated sequences. Pelagibacter-like DNA dominated the vesicle fractions for all samples examined over a depth range of 75 to 500 m. Using a method that provides separation of these three fractions, we compared open ocean depth profiles of DNA associated with each fraction. It is composed of DNA-containing vesicles, viruses, and free DNA and is ubiquitous in all aquatic systems, although the sources, sinks, and ecological consequences are largely unknown. Other potential problems and workarounds with importing Phred results into Sequencher are listed on the Sequencher trouble shooting page.įor more information about Sequencher, please visit more information about using PHRED quality scores in Sequencher,please visit CodonCodeAligner - CodonCodesupport - CodonCodeHome - Phrap.Exocellular DNA is operationally defined as the fraction of the total DNA pool that passes through a membrane filter (0.1 μm). Please note that importing Phrap assemblies into Sequencher can sometimes lead to "holes" in contigs, because Sequencher may clip ends incorrectly when importing Phrap assemblies. In comparison, the MacPHRAP assembly was done more than 40-foldfaster, and led to only two contigs, the larger one covering most ofthe cosmid (44,459 bp).
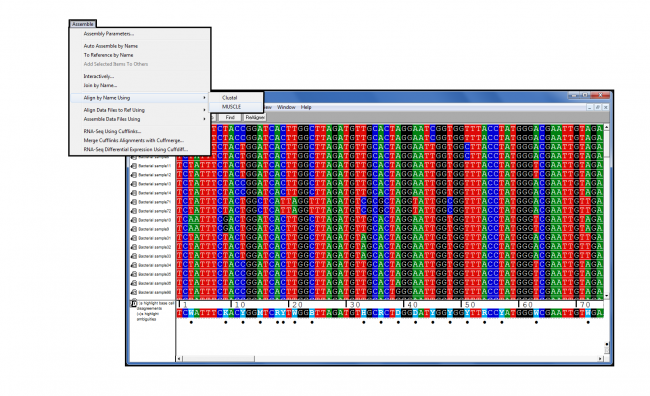
This assembly led tosubstantially fewer (11) and larger (up to 17,386 bp) contigs. Next, the same sequences were imported with PHRED quality scores,and clipped using quality information. This assembly ledto a large number (70) of small (3,305 bp or shorter) contigs.Ģ. In one project, sequence traces without PHRED qualities were used,and ends where clipped using the default settings. Sequencher assemblies were run with default setting and two differentpre-processing steps.ġ. In a test project with more than 800 reads, MacPHRAPassembled the project much faster and into fewer contigs thenSequencher:Ĥ5 minutes 39 seconds (Phred quality scores used) Even just using PHRED-generatedquality scores to perform quality-based end trimming often leads to betterassemblieswith fewer gaps and discrepancies.įor larger sequencing projects with several hundred reads, forexample cosmid or BAC shotgun projects, the combination ofMacPHRED-MacPHRAP with Sequencher 4.1.2 can give faster and betterresults. The newest releases of Sequencher, version4.1.2 and newer, can import PHRAP-generated assemblies. Sequencher is a popular package for Sequence assembly and editingdeveloped by Gene Codes Corporation (no affiliationwith CodonCode Corporation). Gap4 is available for UNIX, Linux, and Windows, and is free toacademic users.For more information about Gap4, please visit the Stadenpackage home page. One potential downside is that it cantake a long time to get familiar with the powerful features Gap4offers. Gap4 andthe preprocessing program Phregap4 offer a number of powerfulinteractive features, as well as options to integrate Phred and Phrapfor base calling and assembly. Gap4 is part of the "Staden package", and was developed by RodgerStaden, James Bonfield, and coworkers at the MRC Cambridge. Please fillout our informationrequest form for pricing information. For more information on obtaining Consed for free for restricted academic use, please visit Consed licenses and support for unrestricted use and use atfor-profit institutions are available through CodonCode. On Mac OSX, Consed requires that X11 (freely available from Apple) is installed.įor academic users and restricted research use, Consed is available for free directly from the authors. Consed is the tool of choice very large projects,or for high-throughput operations that can make use of Consed's "autofinish"features.Ĭonsed is available only for Linux and UNIX, including Mac OS X. You can read more about Aligner, and download demo versions, at ConsedĬonsed was developed specifically for use with Phred and Phrap.Consed offers a interactive functions as well as powerful methods for"autofinishing" - automatic picking of additional sequence reads andwalking primers to close gaps and eliminate low-quality problems inlarge scale sequencing projects.

Phred and Phrap provide functions for base calling and sequenceassembly, but no functions for viewing and editing assemblies.Depending on the operating system you are using, there are severalchoices for "contig editors":ĬodonCode Aligner is a sequence assembly and contig editor program developed with the goal to offer (a) and easy-to-learn, intuitive user interface, and (b) full support for the superior algorithms provided by Phred-Phrap.ĬodonCode Aligner is available for Mac OS X and Windows, and is well suited for small and medium-size projects.
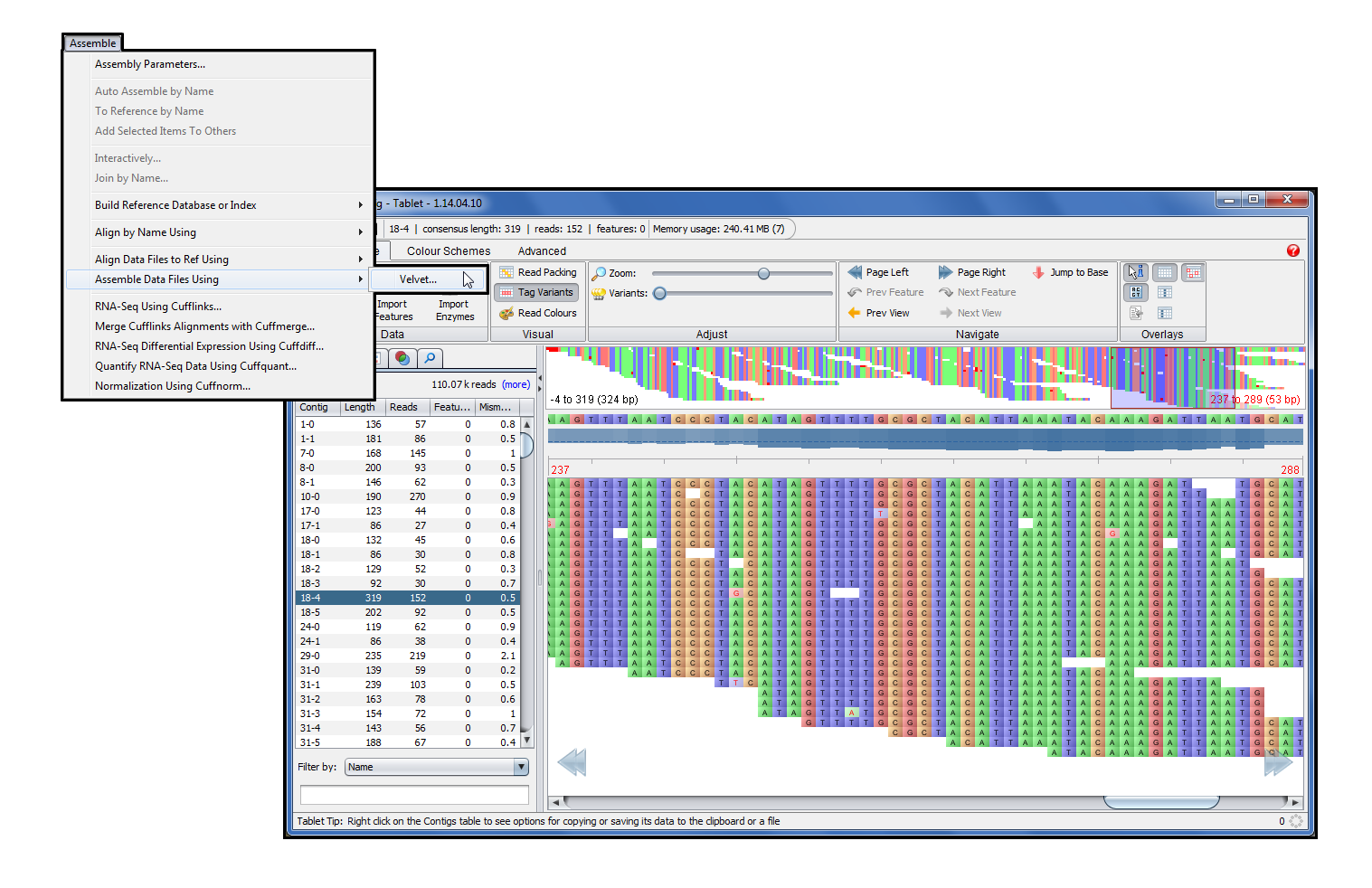
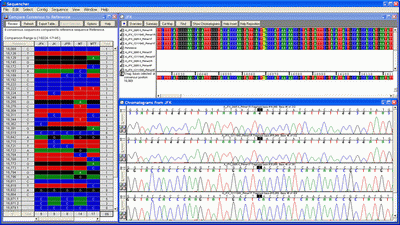
Contig Editors for Phred-Phrap Contig Editors for Phred-Phrap
